We’ll use the gene expression data set described in our previous tutorial: Facilitating Exploratory Data Visualization: Application to TCGA Genomic Data. P2<- insert_yaxis_grob(p1, ydens, grid::unit(.2, "null"), position = "right") P1 <- insert_xaxis_grob(pmain, xdens, grid::unit(.2, "null"), position = "top") Geom_density(data = iris, aes(x = Sepal.Width, fill = Species), Ydens <- axis_canvas(pmain, axis = "y", coord_flip = TRUE)+ 
# Need to set coord_flip = TRUE, if you plan to use coord_flip() Geom_density(data = iris, aes(x = Sepal.Length, fill = Species), Pmain <- ggplot(iris, aes(x = Sepal.Length, y = Sepal.Width, color = Species))+ Recently, in a tweet post, Claus Wilke provides the following solution for creating a perfect scatter plot with marginal density plots or histogram plots: library(cowplot) The problem with the above plots, is the presence of extra spaces between the main plot and the marginal density plots. Yplot <- ggboxplot(iris, x = "Species", y = "Sepal.Width", Xplot <- ggboxplot(iris, x = "Species", y = "Sepal.Length",Ĭolor = "Species", fill = "Species", palette = "jco", # Marginal boxplot of x (top panel) and y (right panel) Size = 3, alpha = 0.6, ggtheme = theme_bw()) Rel_widths = c(2, 1), rel_heights = c(1, 2))Īdd marginal boxplot: # Scatter plot colored by groups ("Species") Plot_grid(xplot, NULL, sp, yplot, ncol = 2, align = "hv", Xplot <- xplot + clean_theme() + rremove("legend") Yplot <- yplot + clean_theme() + rremove("legend") Yplot <- ggdensity(iris, "Sepal.Width", fill = "Species", Xplot <- ggdensity(iris, "Sepal.Length", fill = "Species", # Marginal density plot of x (top panel) and y (right panel) Sp <- ggscatter(iris, x = "Sepal.Length", y = "Sepal.Width", # Scatter plot colored by groups ("Species") In the R code below, we provide a solution using the cowplot package. One limitation of ggExtra is that it can’t cope with multiple groups in the scatter plot and the marginal plots. P <- ggscatter(iris, x = "Sepal.Length", y = "Sepal.Width", The function ggMarginal(), can be used to easily add a marginal histogram, density or boxplot to a scatter plot.įirst, install the ggExtra package as follow: install.packages(“ggExtra”) then type the following R code: # Add density distribution as marginal plot lect: character vector specifying some labels to show.To specify only the size and the style, use font.label = list(size = 14, face = “plain”). For example font.label = list(size = 14, face = “bold”, color =“red”). font.label: a list which can contain the combination of the following elements: the size (e.g.: 14), the style (e.g.: “plain”, “bold”, “italic”, “alic”) and the color (e.g.: “red”) of labels.label: the name of the column containing point labels.By default, geom_smooth() also plots the 95% CI of the best-fit line. We will use the lm method (linear method) plot the best fit line. We will do this by adding geom_smooth() to our ggplot2 figure. Let’s plot the line of best fit (i.e., the line that minimizes the squared difference between the line and each point). This means it is appropriate for us to go ahead and quantify the linear relationship between foot length and subject height. Importantly, there are no unusual data points (e.g., outliers) and the data seem to be distributed relatively linearly (e.g., not u-shaped or exponential). Remember, correlations tell us nothing about causal relationships between variables). People with shorter feet seem to be shorter whereas those with longer feet appear to be taller (or is it the other way round?! People who are shorter have shorter feet whereas those who are taller have longer feet. Scatter_plot + geom_point() + labs(x = "foot length (cm)", y = "height (cm)") Scatter_plot <- ggplot(foot_height, aes(foot, height)) To do so, we need to install the ggplot2 library in R (if not already installed) then load the data into our workspace. Visualizing the relationshipīefore running the correlation analysis, the first thing we need to do is visualize the data. Save the file as indian_foot_height.dat in the working directory of your R session. Right-click on the link and select Save Link As. The dataset we will use contains data on length of the left foot print (col 1) and height (col 2) in 1020 adult male Tamil Indians. In this tutorial we will calculate the correlation between the length of a person’s foot and a person’s height. 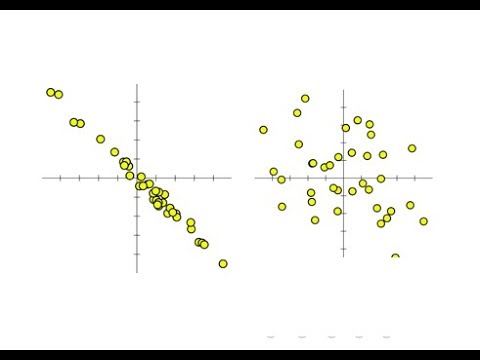
The dataset: foot length and subject height This post assumes you understand the theory behind correlation analysis and have a working knowledge of R it focuses on how to run this type of analysis in R. One simple way to understand and quantify a relationship between two variables is correlation analysis.Īssumptions. Scientists are often interested in understanding the relationship between two variables.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
